Publications
Highlighted
–
A D-2-hydroxyglutarate dehydrogenase mutant reveals a critical role for ketone body metabolism in Caenorhabditis elegans development
PLoS biology
·
12 Apr 2023
·
pmc:PMC10096224
–
All
2024
idh-1neomorphic mutation confers sensitivity to vitamin B12 via increased dependency on one-carbon metabolism inCaenorhabditis elegans
Cold Spring Harbor Laboratory
·
14 Mar 2024
·
doi:10.1101/2024.03.13.584865
2023
Disruption of sugar nucleotide clearance is a therapeutic vulnerability of cancer cells
Nature
·
25 Oct 2023
·
doi:10.1038/s41586-023-06676-3
A D-2-hydroxyglutarate dehydrogenase mutant reveals a critical role for ketone body metabolism in Caenorhabditis elegans development
PLOS Biology
·
12 Apr 2023
·
doi:10.1371/journal.pbio.3002057
A D-2-hydroxyglutarate dehydrogenase mutant reveals a critical role for ketone body metabolism in Caenorhabditis elegans development
PLoS biology
·
12 Apr 2023
·
pmc:PMC10096224
Evolved bacterial resistance to the chemotherapy gemcitabine modulates its efficacy in co-cultured cancer cells
eLife
·
03 Feb 2023
·
doi:10.7554/eLife.83140
2022
Bacterial diet modulates tamoxifen-induced death via host fatty acid metabolism
Nature Communications
·
23 Sep 2022
·
doi:10.1038/s41467-022-33299-5
Teaming up to make kombucha
eLife
·
11 Aug 2022
·
doi:10.7554/eLife.81670
C. elegans as a model for inter-individual variation in metabolism
Nature
·
06 Jul 2022
·
doi:10.1038/s41586-022-04951-3
2021
WormPaths: Caenorhabditis elegans metabolic pathway annotation and visualization
Genetics
·
26 Aug 2021
·
pmc:PMC8864737
WormPaths: Caenorhabditis elegans metabolic pathway annotation and visualization
Genetics
·
12 Jun 2021
·
doi:10.1093/genetics/iyab089
2020
Open collaborative writing with Manubot
PLOS Computational Biology
·
04 Dec 2020
·
doi:10.1371/journal.pcbi.1007128
Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua.
Caenorhabditis elegans methionine/S-adenosylmethionine cycle activity is sensed and adjusted by a nuclear hormone receptor
eLife
·
05 Oct 2020
·
pmc:PMC7561351
Caenorhabditis elegans methionine/S-adenosylmethionine cycle activity is sensed and adjusted by a nuclear hormone receptor
eLife
·
05 Oct 2020
·
doi:10.7554/elife.60259

Constructing knowledge graphs and their biomedical applications
Computational and Structural Biotechnology Journal
·
01 Jan 2020
·
doi:10.1016/j.csbj.2020.05.017
2019
Genome-wide prediction of synthetic rescue mediators of resistance to targeted and immunotherapy
Molecular systems biology
·
11 Mar 2019
·
pmc:PMC6413886
Genome‐wide prediction of synthetic rescue mediators of resistance to targeted and immunotherapy
Molecular Systems Biology
·
01 Mar 2019
·
doi:10.15252/msb.20188323
2018
C. elegans MRP-5 Exports Vitamin B12 from Mother to Offspring to Support Embryonic Development
Cell reports
·
20 Mar 2018
·
pmc:PMC5896776
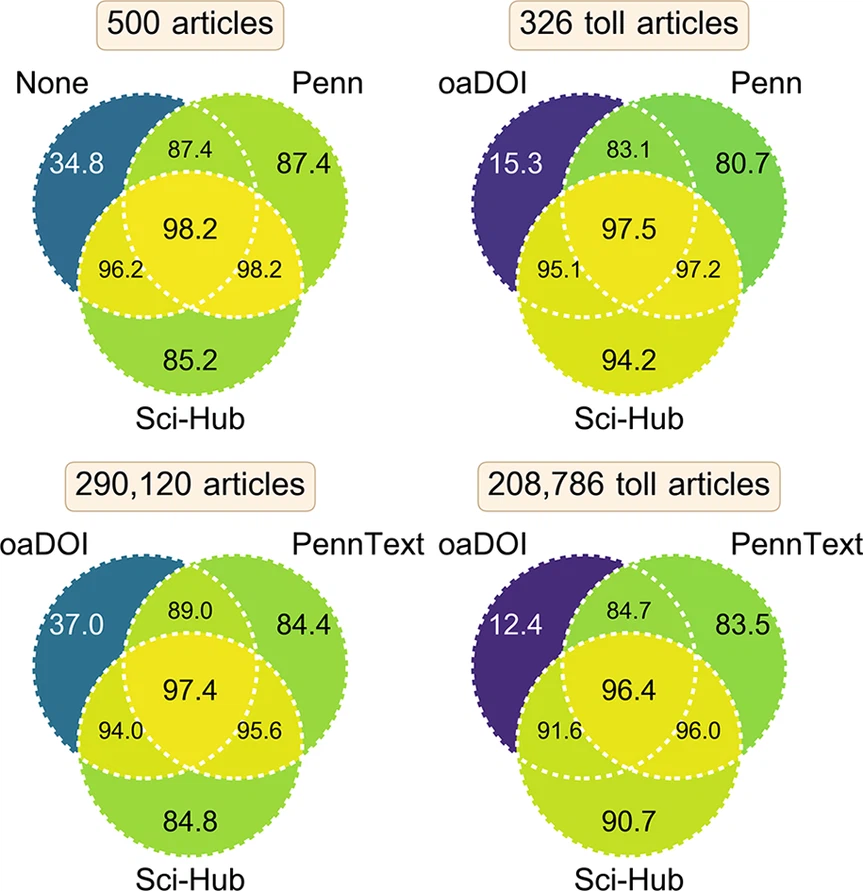
Sci-Hub provides access to nearly all scholarly literature
eLife
·
01 Mar 2018
·
doi:10.7554/eLife.32822
C. elegans MRP-5 Exports Vitamin B12 from Mother to Offspring to Support Embryonic Development
Cell Reports
·
01 Mar 2018
·
doi:10.1016/j.celrep.2018.02.100
2017
Yeast Creates a Niche for Symbiotic Lactic Acid Bacteria through Nitrogen Overflow
Cell systems
·
25 Oct 2017
·
pmc:PMC5660601
Yeast Creates a Niche for Symbiotic Lactic Acid Bacteria through Nitrogen Overflow
Cell Systems
·
01 Oct 2017
·
doi:10.1016/j.cels.2017.09.002
2016
Saccharomyces cerevisiae metabolism in ecological context
FEMS yeast research
·
01 Nov 2016
·
pmc:PMC5050001
Saccharomyces cerevisiaemetabolism in ecological context
FEMS Yeast Research
·
14 Sep 2016
·
doi:10.1093/femsyr/fow080
2015
Metabolic interactions in microbial communities: untangling the Gordian knot
Current Opinion in Microbiology
·
01 Oct 2015
·
doi:10.1016/j.mib.2015.06.014
Metabolic dependencies drive species co-occurrence in diverse microbial communities
Proceedings of the National Academy of Sciences of the United States of America
·
19 May 2015
·
pmc:PMC4443341
Metabolic dependencies drive species co-occurrence in diverse microbial communities
Proceedings of the National Academy of Sciences
·
04 May 2015
·
doi:10.1073/pnas.1421834112
[no title info]
[no publisher info]
·
[no date info]
·
pmid:25941371
[no title info]
[no publisher info]
·
[no date info]
·
pmid:26207681
[no title info]
[no publisher info]
·
[no date info]
·
pmid:27634775
[no title info]
[no publisher info]
·
[no date info]
·
pmid:28964698
[no title info]
[no publisher info]
·
[no date info]
·
pmid:29562169
[no title info]
[no publisher info]
·
[no date info]
·
pmid:30858180
[no title info]
[no publisher info]
·
[no date info]
·
pmid:33016879
[no title info]
[no publisher info]
·
[no date info]
·
pmid:34117752
[no title info]
[no publisher info]
·
[no date info]
·
pmid:37043428